Research
Overview: Addressing Global Food Security
Feeding a growing global population will require major gains in crop productivity under increasingly constrained conditions. By 2050, food production must rise substantially despite declining arable land, increasing climate instability, and persistent disease pressure. In soybean (Glycine max), the soil-borne oomycete Phytophthora sojae remains a major threat, causing stem and root rot and contributing to substantial annual yield losses.
Our research program addresses this challenge by integrating Artificial Intelligence, Functional Genomics, and Microbiology. We aim to build predictive models of how crops interact with pathogens, the soil environment, and the surrounding microbiome to support more resilient and sustainable agriculture.
Research Vision: Decoding Plant Complexity through AI
We study the regulatory logic of plant systems by combining experimental biology with modern computational modeling. Our goal is to transform complex, high-dimensional biological data into mechanistic insight that can guide crop improvement, disease management, and biological discovery.
We use deep learning, graph-based modeling, and multi-omics integration to identify the molecular programs that shape plant health, stress adaptation, and host-pathogen interactions.
Core Research Themes
Our research moves beyond descriptive biology toward causal and predictive understanding through tight integration of in silico modeling and in vivo validation.
1. AI-Driven Genomic Discovery
We develop computational approaches to decode the regulatory grammar of plant genomes. By applying transformer-based architectures and foundation models to genomic and epigenomic data, we identify sequence features and latent patterns associated with gene regulation, stress responses, and adaptive traits.
This work enables us to prioritize candidate regulators and uncover the molecular logic underlying plant resilience.
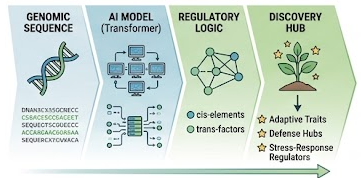
Figure 1. Conceptual overview of the Soy-AI framework, in which genomic sequence is encoded by a foundation model to identify regulatory features, defense hubs, and candidate stress-response regulators.
2. Systems Biology of the Rhizosphere
We investigate the tripartite interaction between the host plant, soil-borne pathogens, and the surrounding microbial community. By integrating transcriptomic, epigenomic, and metagenomic data, we examine the molecular and ecological processes that shape disease outcomes at the root-soil interface.
We use computational pipelines to reconstruct gene regulatory networks (GRNs) and infer how beneficial microbes may suppress disease, stabilize host responses, and promote plant health.
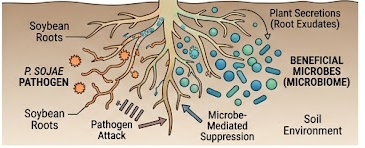
Figure 2. Tripartite interactions in the rhizosphere: soybean roots, beneficial microbes, and soil-borne pathogens collectively shape disease outcome under field stress.
3. Multi-Omics Network Inference
A major focus of the lab is integrating multiple biological layers into unified predictive models. We use graph-based learning and network inference to connect host transcriptional responses, chromatin state, and microbiome variation into interpretable regulatory frameworks.
These models help us identify molecular hubs, cross-talk pathways, and candidate regulators that are most likely to govern plant immunity and stress adaptation.
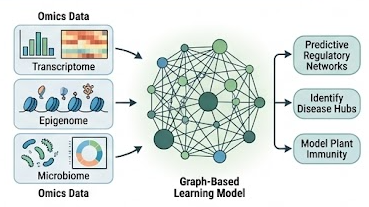
Figure 3. Multi-omics graph modeling integrates transcriptomic, epigenomic, and microbiome signals to reconstruct regulatory networks and identify candidate hubs for plant immunity.
4. Translational Bioinformatics for Sustainability
Our long-term goal is to translate computational insight into practical strategies for crop protection and sustainable agriculture. By understanding the molecular basis of host-pathogen co-evolution and microbiome-mediated resilience, we aim to support:
- more durable genetic resistance
- improved bio-based disease management
- reduced dependence on chemical inputs
- more precise and data-driven agricultural decision-making
This translational perspective connects molecular discovery to field-level resilience.
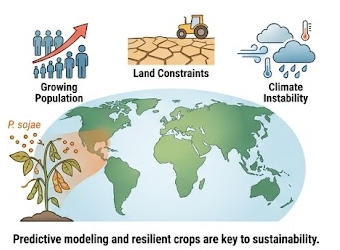
Figure 4. Population growth, land constraints, and disease pressure motivate predictive approaches to crop resilience and sustainable production.
The CompBio Integration
Our research is defined by a tight feedback loop between computation and experimentation. We use computational models to generate high-confidence hypotheses, then test those hypotheses through molecular genetics, microbiology, and plant phenotyping.
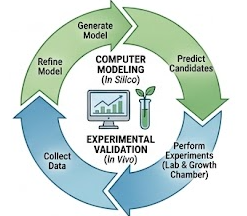
Figure 5. Research workflow linking data collection, AI modeling, hypothesis generation, and experimental validation.
Scalable Computing
We build and maintain high-performance analysis pipelines using Python, R, and HPC environments to process large-scale sequencing and multi-omics datasets, including Nanopore and Illumina platforms.
AI-Ready Biology
We are committed to making complex biological systems machine-learnable. By structuring biological data for predictive modeling, we aim to accelerate discovery and build a foundation for the next generation of AI-enabled plant biology.